Contains models and tools to produce short-term forecasts for both case and sequence notifications assuming circulation of either one or two variants. Tools are also provided to allow the evaluation of the use of sequence data for short-term forecasts in both real-world settings and in user generated scenarios.
Installation
Installing the package
Install the stable development version of the package with:
install.packages("forecast.vocs", repos = "https://epiforecasts.r-universe.dev")Install the unstable development from GitHub using the following,
remotes::install_github("epiforecasts/forecast.vocs", dependencies = TRUE)Installing CmdStan
If you don’t already have CmdStan installed then, in addition to installing forecast.vocs, it is also necessary to install CmdStan using CmdStanR’s install_cmdstan() function to enable model fitting in forecast.vocs. A suitable C++ toolchain is also required. Instructions are provided in the Getting started with CmdStanR vignette. See the CmdStanR documentation for further details and support.
cmdstanr::install_cmdstan()Quick start
This quick start uses data from Germany that includes COVID-19 notifications and sequences with sequences either being positive or negative for the Delta variant. It shows how to produce forecasts for both a one and two strain model for the 19th of June 2021 when the latest available data estimated that approximately 7% of COVID-19 were positive for the Delta variant. Note that estimated growth rates and reproduction numbers shown here have been rescaled using an assumed generation time of 5.5 days and a weakly informative prior centred around Delta being 50% more transmissible than non-Delta cases has been used.
library(forecast.vocs)
options(mc.cores = 4)
obs <- filter_by_availability(
germany_covid19_delta_obs,
date = as.Date("2021-06-19")
)
obs
#> date location_name location cases cases_available seq_total seq_voc
#> 1: 2021-03-20 Germany DE 87328 2021-03-20 NA NA
#> 2: 2021-03-27 Germany DE 109442 2021-03-27 NA NA
#> 3: 2021-04-03 Germany DE 117965 2021-04-03 NA NA
#> 4: 2021-04-10 Germany DE 107223 2021-04-10 NA NA
#> 5: 2021-04-17 Germany DE 142664 2021-04-17 4072 6
#> 6: 2021-04-24 Germany DE 145568 2021-04-24 4500 32
#> 7: 2021-05-01 Germany DE 131887 2021-05-01 3607 55
#> 8: 2021-05-08 Germany DE 107141 2021-05-08 4491 86
#> 9: 2021-05-15 Germany DE 77261 2021-05-15 3443 92
#> 10: 2021-05-22 Germany DE 57310 2021-05-22 3392 114
#> 11: 2021-05-29 Germany DE 33052 2021-05-29 2258 89
#> 12: 2021-06-05 Germany DE 22631 2021-06-05 1136 75
#> 13: 2021-06-12 Germany DE 15553 2021-06-12 NA NA
#> 14: 2021-06-19 Germany DE 7659 2021-06-19 NA NA
#> share_voc seq_available
#> 1: NA <NA>
#> 2: NA <NA>
#> 3: NA <NA>
#> 4: NA <NA>
#> 5: 0.001473477 2021-06-16
#> 6: 0.007111111 2021-06-16
#> 7: 0.015248129 2021-06-16
#> 8: 0.019149410 2021-06-16
#> 9: 0.026720883 2021-06-16
#> 10: 0.033608491 2021-06-16
#> 11: 0.039415412 2021-06-16
#> 12: 0.066021127 2021-06-16
#> 13: NA 2021-07-29
#> 14: NA 2021-07-29
current_obs <- latest_obs(germany_covid19_delta_obs)Forecast all-in-one
Run a forecast for both one and two strain models (or optionally just one of these) using the forecast() function. This provides a wrapper around other package tooling to initialise, fit, and summarise forecasts. Multiple forecasts can be performed efficiently across dates and scenarios using forecast_across_dates() and forecast_across_scenarios().
forecasts <- forecast(obs,
strains = c(1, 2), voc_scale = c(0.4, 0.2),
voc_label = "Delta", scale_r = 5.5 / 7,
adapt_delta = 0.99, max_treedepth = 15,
refresh = 0, show_messages = FALSE
)
forecasts
#> id forecast_date strains overdispersion variant_relationship r_init
#> 1: 0 2021-06-19 1 TRUE correlated 0, 0.25
#> 2: 0 2021-06-19 2 TRUE correlated 0, 0.25
#> voc_scale error fit data fit_args samples max_rhat
#> 1: 0.4, 0.2 <CmdStanMCMC[42]> <list[28]> <list[5]> 4000 1
#> 2: 0.4, 0.2 <CmdStanMCMC[42]> <list[28]> <list[5]> 4000 1
#> divergent_transitions per_divergent_transitions max_treedepth
#> 1: 7 0.00175 9
#> 2: 9 0.00225 10
#> no_at_max_treedepth per_at_max_treedepth time posterior
#> 1: 3693 0.92325 10.6 <fv_posterior[167x20]>
#> 2: 500 0.12500 38.6 <fv_posterior[474x20]>
#> forecast
#> 1: <fv_posterior[12x13]>
#> 2: <fv_posterior[60x13]>Summarised posterior and forecast estimates can be directly extracted from the output using the included summary() method. Alternatively, the complete summarised posterior can be extracted using the following (which also has its own summary() and plot() options to make exploring the results easier). See the documentation for further details.
summary(forecasts, target = "posterior", type = "all")Plot the posterior prediction for cases from the single strain model (“Overall”), from the two strain model (“Combined”), and the unobserved estimates for each strain.
plot(forecasts, current_obs)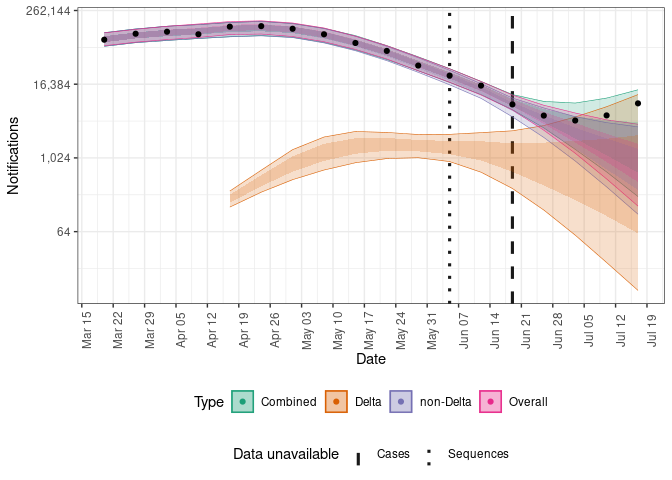
Plot the posterior prediction for the fraction of cases that have the Delta variant from the two strain model.
plot(forecasts, current_obs, type = "voc_frac", voc_label = "Delta variant")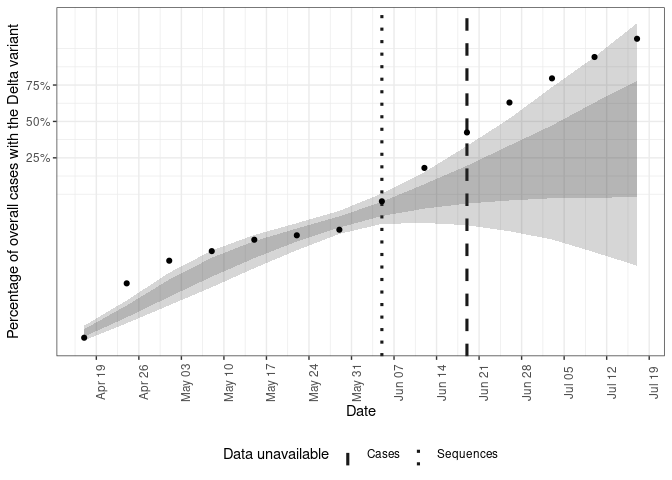
Plot the posterior estimate for the growth rate over the mean of the generation time for COVID-19 cases (here assumed to be 5.5 days).
plot(forecasts, type = "growth")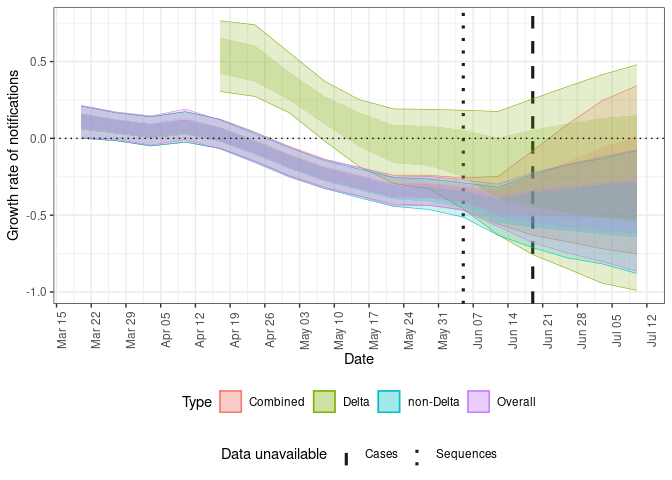
Plot the posterior estimate for the effective reproduction number of Delta and non-Delta cases.
plot(forecasts, type = "rt")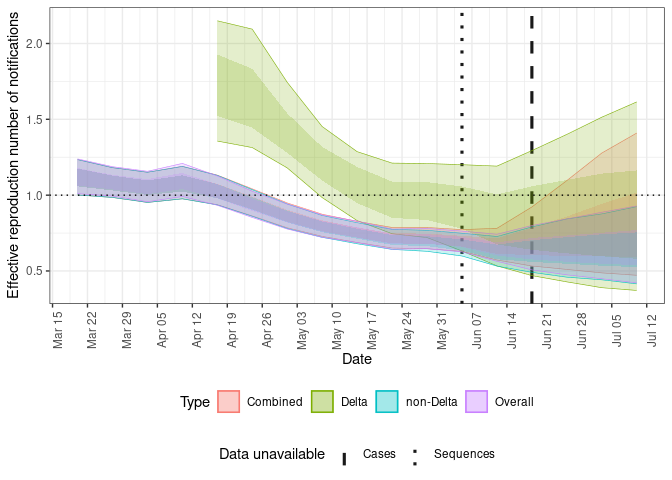
Plot the estimated transmission advantage for Delta vs non-Delta over time. Note that as here a model with a correlated variant growth rates has been used meaning that the difference between variants may vary over time (potentially suitable where imports or present or a variant has greater immune escape than the baseline). If it is likely that the growth advantage is constant over time consider the fixed scaling model. If the variants are likely to be circulating in independent populations consider the independent variant model. See the documentation and model definition vignette for details.
plot(forecasts, type = "voc_advantage")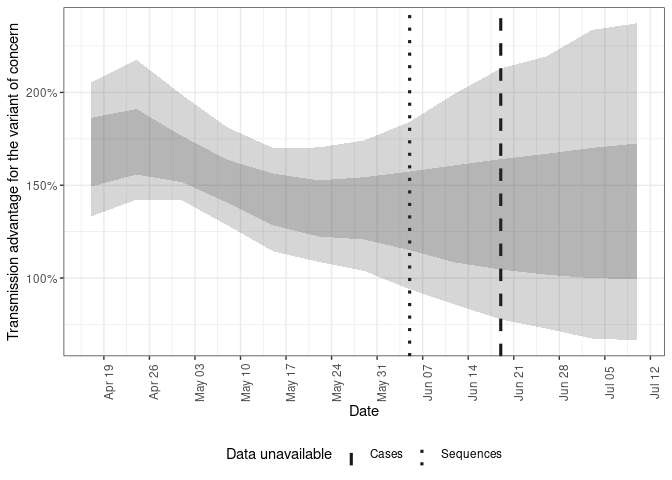
Alternatively a list of all plots with sensible defaults can be produced using the following.
plot(forecasts, current_obs, type = "all", voc_label = "Delta variant")A potentially useful optional integration with scoringutils package is also provided to streamline scoring forecasts using proper scoring rules. See the documentation of the scoringutils package for more on this but as a simple example below we calculate average scores both the single and two strain models for this single Forecast
library(scoringutils)
library(knitr)
summary(forecasts, target = "forecast", type = "cases") |>
fv_score_forecast(current_obs) |>
summarise_scores(by = "strains") |>
kable()| strains | obs_overdispersion | obs_overdispersion | interval_score | dispersion | underprediction | overprediction | coverage_deviation | bias | ae_median |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 1 | 1 | 2056.375 | 277.750 | 1779.750 | 0 | -0.375 | NA | NA |
| 2 | 1 | 1 | 1118.750 | 507.125 | 610.875 | 0 | 0.000 | NA | NA |
Step by step forecast
Rather than using the all-in-one forecast() function individual package functions can be used to produce a forecast as follows.
dt <- fv_as_data_list(obs, horizon = 4)
model <- fv_model(strains = 2)
inits <- fv_inits(dt, strains = 2)
fit <- fv_sample(
data = dt, model = model, init = inits,
voc_scale = c(0.4, 0.2),
adapt_delta = 0.99, max_treedepth = 15,
refresh = 0, show_messages = FALSE
)
posterior <- fv_tidy_posterior(fit, voc_label = "Delta", scale_r = 5.5 / 7)As for forecasts produced with the forecast() function summary plots and estimates can then be produced using summary(), and plot() methods. For example case forecasts can be returned using the following,
summary(posterior, type = "cases", forecast = TRUE)Citation
If using forecast.vocs in your work please consider citing it using the following,
#>
#> To cite forecast.vocs in publications use:
#>
#> Sam Abbott (2021). forecast.vocs: Forecast case and sequence
#> notifications using variant of concern strain dynamics, DOI:
#> 10.5281/zenodo.5559016
#>
#> A BibTeX entry for LaTeX users is
#>
#> @Article{,
#> title = {forecast.vocs: Forecast case and sequence notifications using variant of concern strain dynamics},
#> author = {Sam Abbott},
#> journal = {Zenodo},
#> year = {2021},
#> doi = {10.5281/zenodo.5559016},
#> }How to make a bug report or feature request
Please briefly describe your problem and what output you expect in an issue. If you have a question, please don’t open an issue. Instead, ask on our Q and A page.
Contributing
We welcome contributions and new contributors! We particularly appreciate help on priority problems in the issues. Please check and add to the issues, and/or add a pull request.
Code of Conduct
Please note that the forecast.vocs project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.

